Genetic history of the Iberian Peninsula
The ancestry of modern Iberians (Spanish and Portuguese) is consistent with the geographical situation of the Iberian Peninsula in the south-west corner of Europe. The strongest prehistoric connection is with Atlantic Europe as suggested by the predominance of Y-Chromosome Haplogroup R1b throughout the peninsula. There is also a strong connection with the Eastern Mediterranean region. However, this is lesser than that of the Southeast of the European continent (i.e. the Balkans and Southern Italy) due to Iberia being the farthest away from the Bosphorus, the main bridge of population expansions from Anatolia into Europe during prehistoric times. Indeed, studies point to Eastern Mediterranean genetic contribution to Iberia to have been driven primarily by historical rather than prehistorical population movements (i.e. Phoenicians, Carthaginians, Jews and Levantine Arabs rather than earlier Neolithic farmers). Iberia stands out among other southern European populations as having the highest levels of ancestry originating both in North Africa as well as in Sub-Saharan Africa which is largely ascribed to the long Islamic presence in the Iberian peninsula and possibly African slavery.[1] Significant genetic differences are found among, and even within, Spain's different regions, which can be explained by the wide divergence in their historical trajectories and Spain's internal geographic boundaries. The Basque Region in Northern Spain, is the most genetically distinct and typically Atlantic European. Furthermore, the Basque region and Catalonia hold the least Eastern Mediterranean ancestry in Iberia. African influence is mainly concentrated in the Southern and Western regions of the peninsula.
Population Genetics: Methods and Limitations
One of the first scholars to perform genetic studies was Luigi Luca Cavalli-Sforza. He used classical genetic markers to analyse DNA by proxy. This method studies differences in the frequencies of particular allelic traits, namely polymorphisms from proteins found within human blood (such as the ABO blood groups, Rhesus blood antigens, HLA loci, immunoglobulins, G-6-P-D isoenzymes, among others). Subsequently, his team calculated genetic distance between populations, based on the principle that two populations that share similar frequencies of a trait are more closely related than populations that have more divergent frequencies of the trait.[2]
Since then, population genetics has progressed significantly and studies using direct DNA analysis are now abundant and may use mitochondrial DNA (mtDNA), the non-recombining portion of the Y chromosome (NRY) or autosomal DNA. MtDNA and NRY DNA share some similar features which have made them particularly useful in genetic anthropology. These properties include the direct, unaltered inheritance of mtDNA and NRY DNA from mother to offspring and father to son, respectively, without the 'scrambling' effects of genetic recombination. We also presume that these genetic loci are not affected by natural selection and that the major process responsible for changes in base pairs has been mutation (which can be calculated).[3]
Whereas Y-DNA and mtDNA haplogroups represent but a small component of a person’s DNA pool, autosomal DNA has the advantage of containing hundreds and thousands of examinable genetic loci, thus giving a more complete picture of genetic composition. Descent relationships can only to be determined on a statistical basis, because autosomal DNA undergoes recombination. A single chromosome can record a history for each gene. Autosomal studies are much more reliable for showing the relationships between existing populations but do not offer the possibilities for unraveling their histories in the same way as mtDNA and NRY DNA studies promise, despite their many complications.
Genetic studies operate on numerous assumptions and suffer from methodological limitations such as selection bias and confounding. Phenomenon like genetic drift, foundation and bottleneck effects cause large errors, particularly in haplogroup studies. No matter how accurate the methodology, conclusions derived from such studies are compiled on the basis of how the author envisages their data fits with established archaeological or linguistic theories.
Main genetic compositions

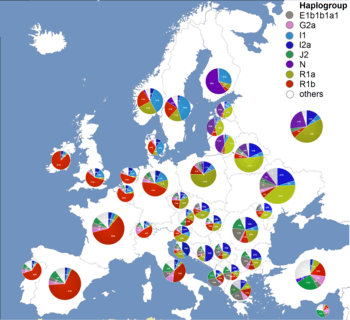
DNA analysis shows that Spanish and Portuguese populations are most closely related to other populations of western Europe.[4][5][6] There is an axis of significant genetic differentiation along the east-west direction, in contrast to remarkable genetic similarity in the north-south direction. North African admixture, associated with the Islamic conquest, can be dated to the period between c. AD 860–1120.[7]
Y-Chromosome haplogroups
Y-DNA Haplogroup R1b is the most frequent among Spaniards and Portuguese, occurring at over 50% throughout most of Spain.[8] R1b is particularly dominant in the Basque Country and Catalonia, occurring at rate of over 80%. In Iberia, most men with R1b belong to the subclade R-P312 (R1b1a1a2a1a2; as of 2017). The distribution of haplogroups other than R1b varies widely from one region to another.
Although R1b prevails in much of Western Europe, a key difference is found in the prevalence in Iberia of R-DF27 (R1b1a1a2a1a2a). This subclade is found in over 60% of the male population in the Basque Country and 40-48% in Madrid, Alicante, Barcelona, Cantabria, Andalucia, Asturias and Galicia.[9] R-DF27 constitutes much more than the half of the total R1b in the Iberian Peninsula. Subsequent in-migration by members of other haplogroups and subclades of R1b did not affect its overall prevalence, although this falls to only two thirds of the total R1b in Valencia and the coast more generally.[8] R-DF27 is also a significant subclade of R1b in parts of France and Britain. However, it is insignificant in Italy; R-S28/R-U152 (R1b1a1a2a1a2b) is the prevailing subclade of R1b in Italy, Switzerland and parts of France, although it represents less than 5.0% of the male population in Iberia. This underlines the lack of any significant genetic impact on the part of Rome, even though the Latin spoken in the Roman Empire was the most significant, or main, source of the modern Spanish and Portuguese language. R-S28/R-U152 is slightly significant in Seville and Barcelona, at 10-20% of the total population, although it is represented at frequencies of only 3.0% in Cantabria and Santander, 2.0% in Castille and Leon, 6% in Valencia, and under 1% in Andalusia.[8] Sephardic Jews I1 0% I2*/I2a 1% I2 0% Haplogroup R1a 5% R1b 13% G 15% Haplogroup J2 2 25% J*/J1 22% E-M2151b1b 9% T 6% Q 2% [10]
Haplogroup J, mostly subclades of Haplogroup J-M172 (J2), is found at levels of over 20% in some regions, while Haplogroup E has a general frequency of about 10% – albeit with peaks surpassing 30% in certain areas. Overall, E-M78 (E1b1b1a1 in 2017) and E-M81 (E1b1b1b1a in 2017) both constitute about 4.0% each, with a further 1.0% from Haplogroup E-M123 (E1b1b1b2a1) and 1.0% from unknown subclades of E-M96.[11] (E-M81 is widely considered to represent relatively historical migrations from North Africa).
| Region | Sample size | C | E | G | I | J2 | JxJ2 | R1a | R1b | Notes |
|---|---|---|---|---|---|---|---|---|---|---|
| Aragon | 34 | 6% | 0% | 18% | 12% | 0% | 3% | 56% | ||
| Andalusia East | 95 | 4% | 3% | 6% | 9% | 3% | 1% | 72% | ||
| Andalusia West | 73 | 15% | 4% | 5% | 14% | 1% | 4% | 54% | ||
| Asturias | 20 | 15% | 5% | 10% | 15% | 0% | 0% | 50% | ||
| Basques | 116 | 1% | 0% | 8% | 3% | 1% | 0% | 87% | ||
| Castilla La Mancha | 63 | 4% | 10% | 2% | 6% | 2% | 2% | 72% | ||
| Castile North-East | 31 | 9% | 3% | 3% | 3% | 0% | 0% | 77% | ||
| Castile North-West | 100 | 19% | 5% | 3% | 8% | 1% | 2% | 60% | ||
| Catalonia | 80 | >0%[13] | 3% | 6% | 3% | 6% | 0% | 0% | 81% | |
| Extremadura | 52 | 18% | 4% | 10% | 12% | 0% | 0% | 50% |
||
| Galicia | 88 | 17% | 6% | 10% | 7% | 1% | 0% | 57% |
||
| Valencia | 73 | >0%[14] | 10% | 1% | 10% | 5% | 3% | 3% | 64% | |
| Majorca | 62 | 9% | 6% | 8% | 8% | 2% | 0% | 66% |
||
| Menorca | 37 | 19% | 0% | 3% | 3% | 0% | 3% | 73% |
||
| Ibiza | 54 | 8% | 13% | 2% | 4% | 0% | 0% | 57% |
||
| Seville | 155 | 7% | 4% | 12% | 8% | 3% | 1% | 60% |
||
| Huelva | 22 | 14% | 0% | 9% | 14% | 0% | 0% | 59% |
||
| Cadiz | 28 | 4% | 0% | 14% | 14% | 4% | 0% | 51% |
||
| Cordoba | 27 | 11% | 0% | 15% | 15% | 0% | 0% | 56% |
||
| Malaga | 26 | 31% | 4% | 0% | 15% | 0% | 8% | 43% |
||
| Leon | 60 | 10% | 7% | 3% | 5% | 2% | 7% | 62% |
||
| Cantabria | 70 | 13% | 9% | 6% | 3% | 3% | 4% | 58% |
Mitochondrial DNA
There have been a number of studies about the mitochondrial DNA haplogroups (mtDNA) in Europe. In contrast to Y DNA haplogroups, mtDNA haplogroups did not show as much geographical patterning, but were more evenly ubiquitous. Apart from the outlying Sami, all Europeans are characterized by the predominance of haplogroups H, U and T. The lack of observable geographic structuring of mtDNA may be due to socio-cultural factors, namely patrilocality and a lack of polyandry.[15]
The subhaplogroups H1 and H3 have been subject to a more detailed study and would be associated to the Magdalenian expansion from Iberia c. 13,000 years ago:[16]
H1 encompasses an important fraction of Western European mtDNA, reaching its local peak among contemporary Basques (27.8%) and appearing at a high frequency among other Iberians and North Africans. Its frequency is above 10% in many other parts of Europe (France, Sardinia, British Isles, Alps, large portions of Eastern Europe), and above 5% in nearly all the continent. Its subclade H1b is most common in eastern Europe and NW Siberia.[17] So far, the highest frequency of H1 - 61%- has been found among the Tuareg of the Fezzan region in Libya.[18][19]
H3 represents a smaller fraction of European genome than H1 but has a somewhat similar distribution with peak among Basques (13.9%), Galicians (8.3%) and Sardinians (8.5%). Its frequency decreases towards the northeast of the continent, though. Studies have suggested haplogroup H3 is highly protective against AIDS progression.[20]
Autosomal DNA
A 2007 European-wide study including Spanish Basques and Valencian Spaniards, found Iberian populations to cluster the furthest from other continental groups, implying that Iberia holds the most ancient European ancestry. In this study, the most prominent genetic stratification in Europe was found to run from the north to the south-east, while another important axis of differentiation runs east-west across the continent. It also found, despite the differences, that all Europeans are closely related.[21]
North African influence
A number of studies have focused on ascertaining the genetic impact of historical North African population movements into Iberia on the genetic composition of modern Spanish and Portuguese populations. Initial studies pointed to the Straits of Gibraltar acting more as a genetic barrier than a bridge during prehistorical times,[22][23][24] while other studies point to a higher level of recent North African admixture among Iberians than among other European populations,[25][26][27][28][11][29][30] albeit this is as a result of more recent migratory movements, particularly the Moorish invasion of Iberia in the 8th century.
In terms of autosomal DNA, the most recent study regarding African admixture in Iberian populations was conducted in April 2013 by Botigué et al. using genome-wide SNP data for over 2000 individuals, concluding that Spain and Portugal hold significant levels of North African ancestry. Estimates of shared ancestry averaged from 4% in some places to 10% in the general population, the populations of the Canary Islands outputted up to 20% of shared ancestry with north Africans, although the Canary islands are a Spanish exclave located in the African continent, and thus this output is not representative of the Iberian population; these same results did not exceed 2% in other western or southern European populations.[1][31][32] However, contrary to past autosomal studies and to what is inferred from Y-Chromosome and Mitochondrial Haplotype frequencies (see below), it does not detect significant levels of Sub-Saharan ancestry in any European population outside the Canary Islands. Indeed, a prior 2011 autosomal study by Moorjani et al. found Sub-Saharan ancestry throughout Europe at ranges of between 1-4%, "the highest proportion of African ancestry in Europe is in Iberia (Portugal 4.2±0.3% and Spain 1.4±0.3%), consistent with inferences based on mitochondrial DNA and Y chromosomes and the observation by Auton et al. that within Europe, the Southwestern Europeans have the highest haplotype-sharing with North Africans."[25][11][29]
In terms of paternal Y-Chromosome DNA, recent studies coincide in that Iberia has the greatest presence of the typically Northwest African Y-chromosome haplotype marker E-M81 in Europe[26][27] as well as Haplotype Va.[33][28] Estimates of Y-Chromosome ancestry vary, with a 2008 study published in the American Journal of Human Genetics using 1140 samples from throughout the Iberian peninsula, giving a proportion of 10.6% North African ancestry[11][29][30] to the paternal composite of iberians, although maternal composition is almost exclusively peninsular. A similar 2009 study of Y-chromosome with 659 samples from Southern Portugal, 680 from Northern Spain, 37 samples from Andalusia, 915 samples from mainland Italy, and 93 samples from Sicily found significally higher levels of North African male ancestry in Portugal, Spain and Sicily (7.7%, 7.1% and 7.5% respectively) than in Italy (5.7%).[26]
Other studies of the Iberian gene-pool have estimated significantly lower levels of North African Ancestry. According to Bosch et al. 2000 "NW African populations may have contributed 7% of Iberian Y chromosomes".[23] A wide-ranging study by Cruciani et al. 2007, using 6,501 unrelated Y-chromosome samples from 81 populations found that: "Considering both these E-M78 sub-haplogroups (E-V12, E-V22, E-V65) and the E-M81 haplogroup, the contribution of northern African lineages to the entire male gene pool of Iberia (barring Pasiegos), continental Italy and Sicily can be estimated as 5.6 percent, 4.6 percent and 6.6 percent, respectively".[34] A 2007 study estimated the contribution of northern African lineages to the entire male gene pool of Iberia as 5.6%."[35] In general aspects, according to (Bosch et al. 2007) "...the origins of the Iberian Y-chromosome pool may be summarized as follows: 5% recent NW African, 78% Upper Paleolithic and later local derivatives (group IX), and 10% Neolithic" (H58, H71).[36]
Recent Mitochondrial DNA studies coincide in that the Iberian Peninsula holds higher levels of typically North African Haplotype U6,[11][29][37][16] as well as higher frequencies of Sub-Saharan African Haplogroup L in Portugal.[16][38][39][39][40][16] However, high frequencies are largely concentrated in the west and south of the Iberian peninsula and therefore overall frequency is higher in Portugal (6.83%) than in Spain (1.9%) with a mean frequency for the entire peninsula of 3.83%. There is considerable geographic divergence across the peninsula with high frequencies observed for South West Castile (8%), Southern Portugal (18.80%), Central Portugal (10.70%), Western Andalusia (14.6%)[40] and Córdoba (8.30%).[16]
Current debates revolve around whether U6 presence is due to Islamic expansion into the Iberian peninsula or prior population movements[11][29][30] and whether Haplogroup L is linked to the slave trade or prior population movements linked to Islamic expansion. A majority of Haplogroup L lineages in Iberia being North African in origin points to the latter.[16][39][25][41][42] In 2015, Hernández et al. concluded that "the estimated entrance of the North African U6 lineages into Iberia at 10 ky correlates well with other L African clades, indicating that some U6 and L lineages moved together from Africa to Iberia in the Early Holocene while a majority were introduced during historic times."[43]
Portuguese mitochondrial DNA genetic diversity
In a study by Sofia L. Marques, Ana Goios, Ana M. Rocha, Maria João Prata, António Amorim, Leonor Gusmão, Cíntia Alves, and Luis Alvarez. "Portuguese mitochondrial DNA genetic diversity- an update and a phylogenetic revision." FSI Genetics 15 (March 2015): pages 27–32. Excerpts from the Abstract:
"[...] In the case of Portugal, previous population genetics studies have already revealed the general portrait of HVS-I and HVS-II mitochondrial diversity, becoming now important to update and expand the mitochondrial region analysed. Accordingly, a total of 292 complete control region sequences from continental Portugal were obtained, under a stringent experimental design to ensure the quality of data through double sequencing of each target region.* Furthermore, H-specific coding region SNPs were examined to detail haplogroup classification and complete mitogenomes were obtained for all sequences belonging to haplogroups U4 and U5. In general, a typical Western European haplogroup or Atlantic modal haplotype composition was found in mainland Portugal, associated to high level of mitochondrial genetic diversity. Within the country, no signs of substructure were detected. The typing of extra coding region SNPs has provided the refinement or confirmation of the previous classification obtained with EMMA tool in 96% of the cases. Finally, it was also possible to enlarge haplogroup U phylogeny with 28 new U4 and U5 mitogenomes."
The Atlantic modal haplotype (AMH) or haplotype 15 is a Y chromosome haplotype of Y-STR microsatellite variations, associated with the Haplogroup R1b. It was discovered prior to many of the SNPs now used to identify subclades of R1b and references to it can be found in some of the older literature. It corresponds most closely with subclade R1b1a2a1a(1) [L11].
The AMH is the most frequently occurring haplotype amongst human males in Atlantic Europe. It is characterized by the following marker alleles:
- DYS388 12
- DYS390 24
- DYS391 11
- DYS392 13
- DYS393 13
- DYS394 14 (also known as DYS19)
The Atlantic modal haplotype reaches the highest frequencies in the Iberian Peninsula, in Great Britain and Ireland. In Portugal it reaches 70% as a whole, with more than 90% in NW Portugal.
See also
| Wikimedia Commons has media related to Genetic studies on Iberian Peninsula. |
References
- 1 2 Botigué, L. R.; Henn, B. M.; Gravel, S; Maples, B. K.; Gignoux, C. R.; Corona, E; Atzmon, G; Burns, E; Ostrer, H; Flores, C; Bertranpetit, J; Comas, D; Bustamante, C. D. (2013). "Gene flow from North Africa contributes to differential human genetic diversity in southern Europe". Proceedings of the National Academy of Sciences. 110 (29): 11791–11796. Bibcode:2013PNAS..11011791B. doi:10.1073/pnas.1306223110. PMC 3718088. PMID 23733930.
- ↑ Cavalli-Sforza (1993, p. 51)
- ↑ Milisauskas (2002, p. 58)
- ↑ Nelis, Mari; Esko, Tõnu; Mägi, Reedik; Zimprich, Fritz; Zimprich, Alexander; Toncheva, Draga; Karachanak, Sena; Piskácková, Tereza; Balascák, Ivan; Peltonen, L; Jakkula, E; Rehnström, K; Lathrop, M; Heath, S; Galan, P; Schreiber, S; Meitinger, T; Pfeufer, A; Wichmann, HE; Melegh, B; Polgár, N; Toniolo, D; Gasparini, P; d'Adamo, P; Klovins, J; Nikitina-Zake, L; Kucinskas, V; Kasnauskiene, J; Lubinski, J; Debniak, T (2009). Fleischer, Robert C., ed. "Genetic Structure of Europeans: A View from the North–East". PLoS ONE. 4 (5): e5472. Bibcode:2009PLoSO...4.5472N. doi:10.1371/journal.pone.0005472. PMC 2675054. PMID 19424496.
- ↑ Wade, Nicholas (13 August 2008). "The Genetic Map of Europe". The New York Times. Retrieved 17 October 2009.
- ↑ Novembre, John; Johnson, Toby; Bryc, Katarzyna; Kutalik, Zoltán; Boyko, Adam R.; Auton, Adam; Indap, Amit; King, Karen S.; Bergmann, Sven; Nelson, Matthew R.; Stephens, Matthew; Bustamante, Carlos D. (2008). "Genes mirror geography within Europe". Nature. 456 (7218): 98–101. Bibcode:2008Natur.456...98N. doi:10.1038/nature07331. PMC 2735096. PMID 18758442. Lay summary – Gene Expression (31 August 2008).
- ↑ Bycroft, C., et al., "Patterns of genetic differentiation and the footprints of historical migrations in the Iberian Peninsula", bioRxiv 250191 (March 2018), doi:10.1101/250191.
- 1 2 3 Myres, Natalie M.; Rootsi, Siiri; Lin, Alice A.; Järve, Mari; King, Roy J.; Kutuev, Ildus; Cabrera, Vicente M.; Khusnutdinova, Elza K.; Pshenichnov, Andrey; Yunusbayev, Bayazit; Balanovsky, Oleg; Balanovska, Elena; Rudan, Pavao; Baldovic, Marian; Herrera, Rene J.; Chiaroni, Jacques; Di Cristofaro, Julie; Villems, Richard; Kivisild, Toomas; Underhill, Peter A. (1 January 2011). "A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe". European Journal of Human Genetics. 19 (1): 95–101. doi:10.1038/ejhg.2010.146. ISSN 1018-4813. PMC 3039512. PMID 20736979.
- ↑ New clues to the evolutionary history of the main European paternal lineage M269: dissection of the Y-SNP S116 in Atlantic Europe and Iberia
- ↑ http://www.eupedia.com/genetics/spain_portugal_dna.shtml#frequency
- 1 2 3 4 5 6 7 Adams, Susan M.; Bosch, Elena; Balaresque, Patricia L.; Ballereau, Stéphane J.; Lee, Andrew C.; Arroyo, Eduardo; López-Parra, Ana M.; Aler, Mercedes; Grifo, Marina S. Gisbert; Brion, Maria; Carracedo, Angel; Lavinha, João; Martínez-Jarreta, Begoña; Quintana-Murci, Lluis; Picornell, Antònia; Ramon, Misericordia; Skorecki, Karl; Behar, Doron M.; Calafell, Francesc; Jobling, Mark A. (2008). "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula". The American Journal of Human Genetics. 83 (6): 725–36. doi:10.1016/j.ajhg.2008.11.007. PMC 2668061. PMID 19061982. Lay summary – Science News (3 January 2009).
- ↑ Flores, Carlos; Maca-Meyer, Nicole; González, Ana M.; Oefner, Peter J.; Shen, Peidong; Pérez, Jose A.; Rojas, Antonio; Larruga, Jose M.; Underhill, Peter A. (28 July 2004). "Reduced genetic structure of the Iberian peninsula revealed by Y-chromosome analysis: implications for population demography". European Journal of Human Genetics. 12 (10): 855–863. doi:10.1038/sj.ejhg.5201225. ISSN 1018-4813. PMID 15280900.
- ↑ Haplogroup C* (C-M130) has been found among males with the surname Llach and originating from Garrotxa, Catalonia, Spain. It was not found among males with the same surname from other areas, or males with other surnames of Catalan origin (Cognoms Catalans, n.d., Resultat; access 15 September 2015). The Cognoms Catalans project, which researches "genetic surnames" in Catalonia, Valencia and the Balearic Islands, is based at Universitat Pompeu Fabra, Barcelona.
- ↑ C Haplogroup – Y-DNA Classic Chart (21 January 2017).
- ↑ Rosser et al. (2000)
- 1 2 3 4 5 6 Pereira, Luisa; Cunha, Carla; Alves, Cintia; Amorim, Antonio (2005). "African Female Heritage in Iberia: A Reassessment of mtDNA Lineage Distribution in Present Times". Human Biology. 77 (2): 213–29. doi:10.1353/hub.2005.0041. PMID 16201138.
- ↑ Loogväli EL, Roostalu U, Malyarchuk BA, et al. (November 2004). "Disuniting uniformity: a pied cladistic canvas of mtDNA haplogroup H in Eurasia". Molecular Biology and Evolution. 21 (11): 2012–21. doi:10.1093/molbev/msh209. PMID 15254257.
- ↑ Ottoni et al. 2010, "Mitochondrial Haplogroup H1 in North Africa: An Early Holocene Arrival from Iberia", Plosone
- ↑ Ottoni et al., "Table of frequencies of haplogroup H1 in the world", Plosone
- ↑ Hendrickson SL, Hutcheson HB, Ruiz-Pesini E, et al. (November 2008). "Mitochondrial DNA Haplogroups influence AIDS Progression". AIDS. 22 (18): 2429–39. doi:10.1097/QAD.0b013e32831940bb. PMC 2699618. PMID 19005266.
- ↑ Bauchet, M; McEvoy, B; Pearson, LN; Quillen, EE; Sarkisian, T; Hovhannesyan, K; Deka, R; Bradley, DG; Shriver, MD (2007). "Measuring European Population Stratification with Microarray Genotype Data". The American Journal of Human Genetics. 80 (5): 948–56. doi:10.1086/513477. PMC 1852743. PMID 17436249.
- ↑ Dupanloup, I.; Bertorelle, G; Chikhi, L; Barbujani, G (2004). "Estimating the Impact of Prehistoric Admixture on the Genome of Europeans". Molecular Biology and Evolution. 21 (7): 1361–72. doi:10.1093/molbev/msh135. PMID 15044595.
- 1 2 Bosch, Elena; Calafell, Francesc; Comas, David; Oefner, Peter J.; Underhill, Peter A.; Bertranpetit, Jaume (2001). "High-Resolution Analysis of Human Y-Chromosome Variation Shows a Sharp Discontinuity and Limited Gene Flow between Northwestern Africa and the Iberian Peninsula". The American Journal of Human Genetics. 68 (4): 1019–29. doi:10.1086/319521. PMC 1275654. PMID 11254456.
- ↑ Comas, David; Calafell, Francesc; Benchemsi, Noufissa; Helal, Ahmed; Lefranc, Gerard; Stoneking, Mark; Batzer, Mark A.; Bertranpetit, Jaume; Sajantila, Antti (2000). "Alu insertion polymorphisms in NW Africa and the Iberian Peninsula: Evidence for a strong genetic boundary through the Gibraltar Straits". Human Genetics. 107 (4): 312–9. doi:10.1007/s004390000370. PMID 11129330.
- 1 2 3 Moorjani P, Patterson N, Hirschhorn JN, Keinan A, Hao L, Atzmon G, et al. (2011). "The History of African Gene Flow into Southern Europeans, Levantines, and Jews". PLoS Genet. 7 (4): e1001373. doi:10.1371/journal.pgen.1001373. PMC 3080861. PMID 21533020.
- 1 2 3 Capelli, Cristian; Onofri, Valerio; Brisighelli, Francesca; Boschi, Ilaria; Scarnicci, Francesca; Masullo, Mara; Ferri, Gianmarco; Tofanelli, Sergio; Tagliabracci, Adriano; Gusmao, Leonor; Amorim, Antonio; Gatto, Francesco; Kirin, Mirna; Merlitti, Davide; Brion, Maria; Verea, Alejandro Blanco; Romano, Valentino; Cali, Francesco; Pascali, Vincenzo (2009). "Moors and Saracens in Europe: estimating the medieval North African male legacy in southern Europe". European Journal of Human Genetics. 17 (6): 848–52. doi:10.1038/ejhg.2008.258. PMC 2947089. PMID 19156170.
- 1 2 Semino, Ornella; Magri, Chiara; Benuzzi, Giorgia; Lin, Alice A.; Al-Zahery, Nadia; Battaglia, Vincenza; MacCioni, Liliana; Triantaphyllidis, Costas; Shen, Peidong; Oefner, Peter J.; Zhivotovsky, Lev A.; King, Roy; Torroni, Antonio; Cavalli-Sforza, L. Luca; Underhill, Peter A.; Santachiara-Benerecetti, A. Silvana (2004). "Origin, Diffusion, and Differentiation of Y-Chromosome Haplogroups E and J: Inferences on the Neolithization of Europe and Later Migratory Events in the Mediterranean Area". The American Journal of Human Genetics. 74 (5): 1023–34. doi:10.1086/386295. PMC 1181965. PMID 15069642.
- 1 2 Gérard, Nathalie; Berriche, Sala; Aouizérate, Annie; Diéterlen, Florent; Lucotte, Gérard (2006). "North African Berber and Arab Influences in the Western Mediterranean Revealed by Y-Chromosome DNA Haplotypes". Human Biology. 78 (3): 307–16. doi:10.1353/hub.2006.0045. PMID 17216803.
- 1 2 3 4 5 González, Ana M.; Brehm, Antonio; Pérez, José A.; Maca-Meyer, Nicole; Flores, Carlos; Cabrera, Vicente M. (2003). "Mitochondrial DNA affinities at the Atlantic fringe of Europe". American Journal of Physical Anthropology. 120 (4): 391–404. doi:10.1002/ajpa.10168. PMID 12627534.
- 1 2 3 Giacomo, F.; Luca, F.; Popa, L. O.; Akar, N.; Anagnou, N.; Banyko, J.; Brdicka, R.; Barbujani, G.; et al. (2004). "Y chromosomal haplogroup J as a signature of the post-neolithic colonization of Europe". Human Genetics. 115 (5): 357–71. doi:10.1007/s00439-004-1168-9. PMID 15322918.
- ↑ Estimating gene flow from North Africa to southern Europe Archived 2015-04-30 at Archive.is, David Comas, one of the authors of the study
- ↑ "La cifra del 20% sólo se da en Canarias, para el resto del país oscila entre el 10% y 12%", explica Comas.", David Comas, one of the authors of the study, Los españoles somos los europeos con más genes magrebíes, Huffington post, June 2013
- ↑ http://digitalcommons.wayne.edu/humbiol/vol73/iss5/11/
- ↑ Cruciani, F.; La Fratta, R.; Trombetta, B.; Santolamazza, P.; Sellitto, D.; Colomb, E. B.; Dugoujon, J. -M.; Crivellaro, F.; Benincasa, T. (2007). "Tracing Past Human Male Movements in Northern/Eastern Africa and Western Eurasia: New Clues from Y-Chromosomal Haplogroups E-M78 and J-M12". Molecular Biology and Evolution. 24 (6): 1300–1311. doi:10.1093/molbev/msm049. PMID 17351267.
- ↑ Cruciani, F.; La Fratta, R.; Trombetta, B.; Santolamazza, P.; Sellitto, D.; Colomb, E. B.; Dugoujon, J.-M.; Crivellaro, F.; Benincasa, T.; Pascone, R.; Moral, P.; Watson, E.; Melegh, B.; Barbujani, G.; Fuselli, S.; Vona, G.; Zagradisnik, B.; Assum, G.; Brdicka, R.; Kozlov, A. I.; Efremov, G. D.; Coppa, A.; Novelletto, A.; Scozzari, R. (2007). "Tracing Past Human Male Movements in Northern/Eastern Africa and Western Eurasia: New Clues from Y-Chromosomal Haplogroups E-M78 and J-M12". Molecular Biology and Evolution. 24 (6): 1300–11. doi:10.1093/molbev/msm049. PMID 17351267.
- ↑ Bosch, E; Calafell, F; Comas, D; Oefner, PJ; Underhill, PA; Bertranpetit, J (April 2001). "High-resolution analysis of human Y-chromosome variation shows a sharp discontinuity and limited gene flow between northwestern Africa and the Iberian Peninsula". Am. J. Hum. Genet. 68 (4): 1019–29. doi:10.1086/319521. PMC 1275654. PMID 11254456.
- ↑ Plaza, S.; Calafell, F.; Helal, A.; Bouzerna, N.; Lefranc, G.; Bertranpetit, J.; Comas, D. (2003). "Joining the Pillars of Hercules: mtDNA Sequences Show Multidirectional Gene Flow in the Western Mediterranean". Annals of Human Genetics. 67 (Pt 4): 312–28. doi:10.1046/j.1469-1809.2003.00039.x. PMID 12914566.
- ↑ http://genome.cshlp.org/content/22/5/821.full#ref-12
- 1 2 3 Brehm A, Pereira L, Kivisild T, Amorim A (December 2003). "Mitochondrial portraits of the Madeira and Açores archipelagos witness different genetic pools of its settlers". Human Genetics. 114 (1): 77–86. doi:10.1007/s00439-003-1024-3. PMID 14513360.
- 1 2 http://www.biomedcentral.com/content/pdf/1471-2156-15-11.pdf
- ↑ Gonzalez et al. (2003)
- ↑ Alvarez; et al. (2010). "Mitochondrial DNA Patterns in the Iberian Northern Plateau: Population Dynamics and Substructure of the Zamora Province". Am. J. Phys. Anthropol. 142 (4): 531–9. doi:10.1002/ajpa.21252. PMID 20127843.
- ↑ Hernández, Candela L.; Soares, Pedro; Dugoujon, Jean M.; Novelletto, Andrea; Rodríguez, Juan N.; Rito, Teresa; et al. (2015). "Early Holocenic and Historic mtDNA African Signatures in the Iberian Peninsula: Southern Portugal and the Andalusian Region as a Paradigm". PLoS ONE. 10 (10): e0139784. Bibcode:2015PLoSO..1039784H. doi:10.1371/journal.pone.0139784. PMC 4624789. PMID 26509580.