Photolyase
| FAD binding domain of DNA photolyase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
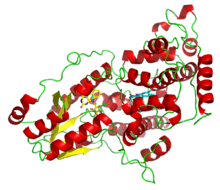 A deazaflavin photolyase from Anacystis nidulans, illustrating the two light-harvesting cofactors: FADH− (yellow) and 8-HDF (cyan). | |||||||||
| Identifiers | |||||||||
| Symbol | FAD_binding_7 | ||||||||
| Pfam | PF03441 | ||||||||
| InterPro | IPR005101 | ||||||||
| PROSITE | PDOC00331 | ||||||||
| SCOP | 1qnf | ||||||||
| SUPERFAMILY | 1qnf | ||||||||
| |||||||||
Photolyases (EC 4.1.99.3) are DNA repair enzymes that repair damage caused by exposure to ultraviolet light. These enzymes require visible light (from the violet/blue end of the spectrum) both for their own activation[1] and for the actual DNA repair.[2] The DNA repair mechanism involving photolyases is called photoreactivation.
Photolyase is a phylogenetically old enzyme which is present and functional in many species, from the bacteria to the fungi to plants[3] and to the animals.[4] Photolyase is particularly important in repairing UV induced damage in plants. The photolyase mechanism is no longer working in humans and other placental mammals who instead rely on the less efficient nucleotide excision repair mechanism.[5]
Photolyases bind complementary DNA strands and break certain types of pyrimidine dimers that arise when a pair of thymine or cytosine bases on the same strand of DNA become covalently linked. The bond length of this dimerization is shorter than the bond length of normal B-DNA structure which produces an incorrect template for replication and transcription.[6] The more common covalent linkage involves the formation of a cyclobutane bridge. Photolyases have a high affinity for these lesions and reversibly bind and convert them back to the original bases.
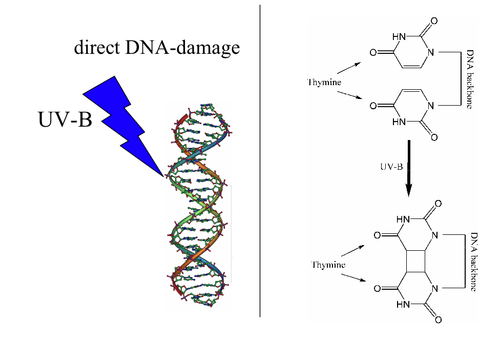
Photolyases are flavoproteins and contain two light-harvesting cofactors. All photolyases contain the two-electron-reduced FADH−; they are divided into two main classes based on the second cofactor, which may be either the pterin methenyltetrahydrofolate (MTHF) in folate photolyases or the deazaflavin 8-hydroxy-7,8-didemethyl-5-deazariboflavin (8-HDF) in deazaflavin photolyases. Although only FAD is required for catalytic activity, the second cofactor significantly accelerates reaction rate in low-light conditions. The enzyme acts by electron transfer in which the reduced flavin FADH− is activated by light energy and acts as an electron donor to break the pyrimidine dimer.[7]
On the basis of sequence similarities DNA photolyases can be grouped into two classes. The first class contains enzymes from Gram-negative and Gram-positive bacteria, the halophilic archaea Halobacterium halobium, fungi and plants. Proteins containing this domain also include Arabidopsis thaliana cryptochromes 1 and 2, which are blue light photoreceptors that mediate blue light-induced gene expression and modulation of circadian rhythms.
The second class are named cryptochrome (Cry), found in species as diverse as Drosophila, Arabidopsis, Synechocystis, and Human (Cry-DASH). These were previously assumed to have no DNA repair activity because of negligible activity on double-stranded DNA. A study[4] by A. Sancar and P. Selby provided evidence to suggest this branch of cryptochromes have photolyase activity with a high degree of specificity for cyclobutane pyrimidine dimers in single-stranded DNA. Their study showed that VcCry1 from Vibrio cholerae, X1Cry from Xenopus laevis, and AtCry3 from Arabidopsis thaliana all had photolyase activity on UV irradiated ssDNA in vitro.
Some sunscreens include photolyase in their ingredients, claiming a reparative action on UV-damaged skin.[8]
Human proteins containing this domain
References
- ↑ Yamamoto J, Shimizu K, Kanda T, Hosokawa Y, Iwai S, Plaza P, Müller P (October 2017). "Loss of Fourth Electron-Transferring Tryptophan in Animal (6-4) Photolyase Impairs DNA Repair Activity in Bacterial Cells". Biochemistry. 56 (40): 5356–5364. doi:10.1021/acs.biochem.7b00366. PMID 28880077.
- ↑ Thiagarajan V, Byrdin M, Eker AP, Müller P, Brettel K (June 2011). "Kinetics of cyclobutane thymine dimer splitting by DNA photolyase directly monitored in the UV". Proceedings of the National Academy of Sciences of the United States of America. 108 (23): 9402–7. doi:10.1073/pnas.1101026108. PMC 3111307. PMID 21606324.
- ↑ Teranishi, M., Nakamura, K., Morioka, H.,Yamamoto, K. and Hidema, J. (2008). "The native cyclobutane pyrimidine dimer photolyase of rice is phosphorylated". Plant Physiology. 146 (4): 1941–1951. doi:10.1104/pp.107.110189. PMC 2287361. PMID 18235036.
- 1 2 Selby CP, Sancar A (November 2006). "A cryptochrome/photolyase class of enzymes with single-stranded DNA-specific photolyase activity". Proceedings of the National Academy of Sciences of the United States of America. 103 (47): 17696–700. doi:10.1073/pnas.0607993103. PMC 1621107. PMID 17062752.
- ↑ Lucas-Lledó JI, Lynch M (May 2009). "Evolution of mutation rates: phylogenomic analysis of the photolyase/cryptochrome family". Molecular Biology and Evolution. 26 (5): 1143–53. doi:10.1093/molbev/msp029. PMC 2668831. PMID 19228922.
- ↑ Garrett RH, Grisham CM (2010). Biochemistry. Brooks/Cole, Cengage Learning. ISBN 978-0-495-10935-8. OCLC 984382855.
- ↑ Sancar A (June 2003). "Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors". Chemical Reviews. 103 (6): 2203–37. doi:10.1021/cr0204348. PMID 12797829.
- ↑ Kulms D, Pöppelmann B, Yarosh D, Luger TA, Krutmann J, Schwarz T (July 1999). "Nuclear and cell membrane effects contribute independently to the induction of apoptosis in human cells exposed to UVB radiation". Proceedings of the National Academy of Sciences of the United States of America. 96 (14): 7974–9. doi:10.1073/pnas.96.14.7974. PMC 22172. PMID 10393932.