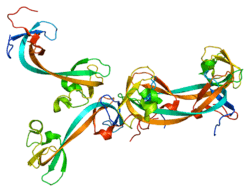KDM4A
Lysine-specific demethylase 4A is an enzyme that in humans is encoded by the KDM4A gene.[5][6][7]
Function
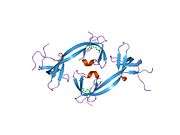
This gene is a member of the Jumonji domain 2 (JMJD2) family and encodes a protein with a JmjN domain, a JmjC domain, a JD2H domain, two TUDOR domains, and two PHD-type zinc fingers. This nuclear protein belongs to the alpha-ketoglutarate-dependent hydroxylase superfamily. It functions as a trimethylation-specific demethylase, converting specific trimethylated histone on histone H3 lysine 9 and 36 residues to the dimethylated form and lysine 9 dimethylated residues to monomethyl, and as a transcriptional repressor.[7]
Alterations in this gene have been found associated with chromosomal instability that leads to cancer.( PMID 23871696)
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000066135 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000033326 - Ensembl, May 2017
- ↑ "Human PubMed Reference:".
- ↑ "Mouse PubMed Reference:".
- ↑ Ishikawa K, Nagase T, Suyama M, Miyajima N, Tanaka A, Kotani H, Nomura N, Ohara O (June 1998). "Prediction of the coding sequences of unidentified human genes. X. The complete sequences of 100 new cDNA clones from brain which can code for large proteins in vitro". DNA Research. 5 (3): 169–76. doi:10.1093/dnares/5.3.169. PMID 9734811.
- ↑ Katoh M, Katoh M (June 2004). "Identification and characterization of JMJD2 family genes in silico". International Journal of Oncology. 24 (6): 1623–8. doi:10.3892/ijo.25.3.759. PMID 15138608.
- 1 2 "Entrez Gene: JMJD2A jumonji domain containing 2A".
Further reading
- Bonaldo MF, Lennon G, Soares MB (September 1996). "Normalization and subtraction: two approaches to facilitate gene discovery". Genome Research. 6 (9): 791–806. doi:10.1101/gr.6.9.791. PMID 8889548.
- Yoon HG, Chan DW, Reynolds AB, Qin J, Wong J (September 2003). "N-CoR mediates DNA methylation-dependent repression through a methyl CpG binding protein Kaiso". Molecular Cell. 12 (3): 723–34. doi:10.1016/j.molcel.2003.08.008. PMID 14527417.
- Brandenberger R, Wei H, Zhang S, Lei S, Murage J, Fisk GJ, Li Y, Xu C, Fang R, Guegler K, Rao MS, Mandalam R, Lebkowski J, Stanton LW (June 2004). "Transcriptome characterization elucidates signaling networks that control human ES cell growth and differentiation". Nature Biotechnology. 22 (6): 707–16. doi:10.1038/nbt971. PMID 15146197.
- Suzuki Y, Yamashita R, Shirota M, Sakakibara Y, Chiba J, Mizushima-Sugano J, Nakai K, Sugano S (September 2004). "Sequence comparison of human and mouse genes reveals a homologous block structure in the promoter regions". Genome Research. 14 (9): 1711–8. doi:10.1101/gr.2435604. PMC 515316. PMID 15342556.
- Gray SG, Iglesias AH, Lizcano F, Villanueva R, Camelo S, Jingu H, Teh BT, Koibuchi N, Chin WW, Kokkotou E, Dangond F (August 2005). "Functional characterization of JMJD2A, a histone deacetylase- and retinoblastoma-binding protein". The Journal of Biological Chemistry. 280 (31): 28507–18. doi:10.1074/jbc.M413687200. PMID 15927959.
- Zhang D, Yoon HG, Wong J (August 2005). "JMJD2A is a novel N-CoR-interacting protein and is involved in repression of the human transcription factor achaete scute-like homologue 2 (ASCL2/Hash2)". Molecular and Cellular Biology. 25 (15): 6404–14. doi:10.1128/MCB.25.15.6404-6414.2005. PMC 1190321. PMID 16024779.
- Tao WA, Wollscheid B, O'Brien R, Eng JK, Li XJ, Bodenmiller B, Watts JD, Hood L, Aebersold R (August 2005). "Quantitative phosphoproteome analysis using a dendrimer conjugation chemistry and tandem mass spectrometry". Nature Methods. 2 (8): 591–8. doi:10.1038/nmeth776. PMID 16094384.
- Kim J, Daniel J, Espejo A, Lake A, Krishna M, Xia L, Zhang Y, Bedford MT (April 2006). "Tudor, MBT and chromo domains gauge the degree of lysine methylation". EMBO Reports. 7 (4): 397–403. doi:10.1038/sj.embor.7400625. PMC 1456902. PMID 16415788.
- Huang Y, Fang J, Bedford MT, Zhang Y, Xu RM (May 2006). "Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A". Science. 312 (5774): 748–51. doi:10.1126/science.1125162. PMID 16601153.
- Whetstine JR, Nottke A, Lan F, Huarte M, Smolikov S, Chen Z, Spooner E, Li E, Zhang G, Colaiacovo M, Shi Y (May 2006). "Reversal of histone lysine trimethylation by the JMJD2 family of histone demethylases". Cell. 125 (3): 467–81. doi:10.1016/j.cell.2006.03.028. PMID 16603238.
- Chen Z, Zang J, Whetstine J, Hong X, Davrazou F, Kutateladze TG, Simpson M, Mao Q, Pan CH, Dai S, Hagman J, Hansen K, Shi Y, Zhang G (May 2006). "Structural insights into histone demethylation by JMJD2 family members". Cell. 125 (4): 691–702. doi:10.1016/j.cell.2006.04.024. PMID 16677698.
- Klose RJ, Yamane K, Bae Y, Zhang D, Erdjument-Bromage H, Tempst P, Wong J, Zhang Y (July 2006). "The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine 9 and lysine 36". Nature. 442 (7100): 312–6. doi:10.1038/nature04853. PMID 16732292.
- Shin S, Janknecht R (August 2007). "Activation of androgen receptor by histone demethylases JMJD2A and JMJD2D". Biochemical and Biophysical Research Communications. 359 (3): 742–6. doi:10.1016/j.bbrc.2007.05.179. PMID 17555712.
- Chen Z, Zang J, Kappler J, Hong X, Crawford F, Wang Q, Lan F, Jiang C, Whetstine J, Dai S, Hansen K, Shi Y, Zhang G (June 2007). "Structural basis of the recognition of a methylated histone tail by JMJD2A". Proceedings of the National Academy of Sciences of the United States of America. 104 (26): 10818–23. doi:10.1073/pnas.0704525104. PMC 1891149. PMID 17567753.
- Ng SS, Kavanagh KL, McDonough MA, Butler D, Pilka ES, Lienard BM, Bray JE, Savitsky P, Gileadi O, von Delft F, Rose NR, Offer J, Scheinost JC, Borowski T, Sundstrom M, Schofield CJ, Oppermann U (July 2007). "Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity". Nature. 448 (7149): 87–91. doi:10.1038/nature05971. PMID 17589501.
- Lee J, Thompson JR, Botuyan MV, Mer G (January 2008). "Distinct binding modes specify the recognition of methylated histones H3K4 and H4K20 by JMJD2A-tudor". Nature Structural & Molecular Biology. 15 (1): 109–11. doi:10.1038/nsmb1326. PMC 2211384. PMID 18084306.
This article is issued from
Wikipedia.
The text is licensed under Creative Commons - Attribution - Sharealike.
Additional terms may apply for the media files.