MAP1LC3B
Microtubule-associated proteins 1A/1B light chain 3B (hereafter referred to as LC3) is a protein that in humans is encoded by the MAP1LC3B gene.[5] LC3 is a central protein in the autophagy pathway where it functions in autophagy substrate selection and autophagosome biogenesis. LC3 is the most widely used marker of autophagosomes.[6]
Discovery
LC3 was originally identified as a microtubule associated protein in rat brain.[7] However it was later found that the primary function of LC3 is in autophagy, a process that involves the bulk degradation of cytoplasmic components.
The ATG8 protein family
MAP1LC3B is a member of the highly conserved ATG8 protein family. ATG8 proteins are present in all known eukaryotic organisms. The animal ATG8 family comprises three subfamilies: (i) microtubule-associated protein 1 light chain 3 (MAP1LC3); (ii) Golgi-associated ATPase enhancer of 16 kDa (GATE-16); and (iii) γ-amino-butyric acid receptor-associate protein (GABARAP). MAP1LC3B is one of the four genes in the MAP1LC3 subfamily (others include MAP1LC3A, MAP1LC3C, and MAP1LC3B2).[8]
Function
Cytoplasmic LC3
Newly synthesized LC3's C-terminus is hydroylzed by a cysteine protease called ATG4B exposing Gly120, termed LC3-I.[9] LC3-I, through a series of ubiquitin-like reactions involving enzymes ATG7, ATG3, and ATG12-ATG5-ATG16, becomes conjugated to the head group of the lipid phosphatidylethanolamine.[10] The lipid modified form of LC3, referred to as LC3-II, is believed to be involved in autophagosome membrane expansion and fusion events.[11] However, the exact role of LC3 in the autophagic pathway is still discussed. And the question of whether LC3 is required for autophagy is debated since knockdown of MAP1LC3B is compensated by the other members of the MAP1LC3 subfamily. Previous studies showed that MAP1LC3B knock out mice develop normally, possibly due to a then unknown compensatory mechanism.[12] Further work, however, demonstrated that LC3 is required for autophagy by simultaneously down-regulating all of the MAP1LC3 subfamily members.[13] While yet another study argues that MAP1LC3 knockdown does to not affect bulk autophagy, whereas its GABARAP family members are crucial for the process.[14][14] LC3 also functions—together with autophagy receptors (e.g. SQSTM1)--in the selective capture of cargo for autophagic degradation.[15] Independent of autophagosomes, a single soluble LC3 is associated with an approximately 500 kDa complex in the cytoplasm.[16]
Nuclear LC3
The importance of the nuclear functions of autophagy proteins should not be underestimated. A large pool of LC3 is present in the nucleus of a variety of different cell types.[17] In response to starvation, nuclear LC3 is deacetylated and trafficked out of the nucleus into the cytoplasm where it functions in autophagy.[18] Nuclear LC3 interacts with lamin B1, and participates in the degradation of nuclear lamina.[19] LC3 is also enriched in nucleoli via its triple arginine motif, and associates with a number of different nuclear and nucleolar constituents including: MAP1B, tubulin, and several ribosomal proteins.[20]
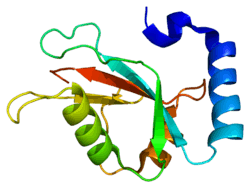
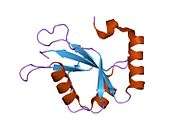
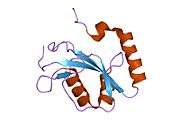
Structure
LC3 shares structural homology with ubiquitin, and thus has been termed a ubiquitin-like protein.[21] LC3 has a LDS (LIR docking site)/hydrophobic binding interface in the N terminus which interacts with LIR (LC3 Interacting Region) containing proteins.[16] This domain is rich in hydrophobic amino acids, the mutation of which impairs the ability of LC3 binding with LIR containing proteins, many of which are autophagy cargo adapter proteins. For example, sequestosome (SQSTM1) interacts with Phe 52 and Leu53 aminoacids present in hydrophobic binding interface of LC3 and any mutation of these amino acids prevents LC3 interaction with SQSTM1.
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000140941 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000031812 - Ensembl, May 2017
- ↑ "Human PubMed Reference:".
- ↑ "Mouse PubMed Reference:".
- ↑ "Entrez Gene: MAP1LC3B microtubule-associated protein 1 light chain 3 beta".
- ↑ Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, et al. (Jan 2016). "Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition)". Autophagy. 12 (1): 1–222. doi:10.1080/15548627.2015.1100356. PMID 26799652.
- ↑ Mann SS, Hammarback JA (April 1994). "Molecular characterization of light chain 3. A microtubule binding subunit of MAP1A and MAP1B". The Journal of Biological Chemistry. 269 (15): 11492–7. PMID 7908909.
- ↑ Shpilka T, Weidberg H, Pietrokovski S, Elazar Z (July 2011). "Atg8: an autophagy-related ubiquitin-like protein family". Genome Biology. 12 (7): 226. doi:10.1186/gb-2011-12-7-226. PMC 3218822. PMID 21867568.
- ↑ Kirisako T, Ichimura Y, Okada H, Kabeya Y, Mizushima N, Yoshimori T, Ohsumi M, Takao T, Noda T, Ohsumi Y (October 2000). "The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway". The Journal of Cell Biology. 151 (2): 263–76. doi:10.1083/jcb.151.2.263. PMC 2192639. PMID 11038174.
- ↑ Ohsumi Y (March 2001). "Molecular dissection of autophagy: two ubiquitin-like systems". Nature Reviews. Molecular Cell Biology. 2 (3): 211–6. doi:10.1038/35056522. PMID 11265251.
- ↑ Weidberg H, Shpilka T, Shvets E, Abada A, Shimron F, Elazar Z (April 2011). "LC3 and GATE-16 N termini mediate membrane fusion processes required for autophagosome biogenesis". Developmental Cell. 20 (4): 444–54. doi:10.1016/j.devcel.2011.02.006. PMID 21497758.
- ↑ Cann GM, Guignabert C, Ying L, Deshpande N, Bekker JM, Wang L, Zhou B, Rabinovitch M (January 2008). "Developmental expression of LC3alpha and beta: absence of fibronectin or autophagy phenotype in LC3beta knockout mice". Developmental Dynamics. 237 (1): 187–95. doi:10.1002/dvdy.21392. PMID 18069693.
- ↑ Weidberg H, Shvets E, Shpilka T, Shimron F, Shinder V, Elazar Z (June 2010). "LC3 and GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis". The EMBO Journal. 29 (11): 1792–802. doi:10.1038/emboj.2010.74. PMC 2885923. PMID 20418806.
- 1 2 Szalai P, Hagen LK, Sætre F, Luhr M, Sponheim M, Øverbye A, Mills IG, Seglen PO, Engedal N (April 2015). "Autophagic bulk sequestration of cytosolic cargo is independent of LC3, but requires GABARAPs". Experimental Cell Research. 333 (1): 21–38. doi:10.1016/j.yexcr.2015.02.003. PMID 25684710.
- ↑ Johansen T, Lamark T (March 2011). "Selective autophagy mediated by autophagic adapter proteins". Autophagy. 7 (3): 279–96. doi:10.4161/auto.7.3.14487. PMC 3060413. PMID 21189453.
- 1 2 Kraft LJ, Nguyen TA, Vogel SS, Kenworthy AK (May 2014). "Size, stoichiometry, and organization of soluble LC3-associated complexes". Autophagy. 10 (5): 861–77. doi:10.4161/auto.28175. PMC 4768459. PMID 24646892.
- ↑ Drake KR, Kang M, Kenworthy AK (March 2010). "Nucleocytoplasmic distribution and dynamics of the autophagosome marker EGFP-LC3". PLoS One. 5 (3): e9806. doi:10.1371/journal.pone.0009806. PMC 2843706. PMID 20352102.
- ↑ Huang R, Xu Y, Wan W, Shou X, Qian J, You Z, Liu B, Chang C, Zhou T, Lippincott-Schwartz J, Liu W (February 2015). "Deacetylation of nuclear LC3 drives autophagy initiation under starvation". Molecular Cell. 57 (3): 456–66. doi:10.1016/j.molcel.2014.12.013. PMID 25601754.
- ↑ Dou Z, Xu C, Donahue G, Shimi T, Pan JA, Zhu J, Ivanov A, Capell BC, Drake AM, Shah PP, Catanzaro JM, Ricketts MD, Lamark T, Adam SA, Marmorstein R, Zong WX, Johansen T, Goldman RD, Adams PD, Berger SL (November 2015). "Autophagy mediates degradation of nuclear lamina". Nature. 527 (7576): 105–9. doi:10.1038/nature15548. PMC 4824414. PMID 26524528.
- ↑ Kraft LJ, Manral P, Dowler J, Kenworthy AK (April 2016). "Nuclear LC3 Associates with Slowly Diffusing Complexes that Survey the Nucleolus". Traffic. 17 (4): 369–99. doi:10.1111/tra.12372. PMC 4975375. PMID 26728248.
- ↑ Kouno T, Mizuguchi M, Tanida I, Ueno T, Kanematsu T, Mori Y, Shinoda H, Hirata M, Kominami E, Kawano K (July 2005). "Solution structure of microtubule-associated protein light chain 3 and identification of its functional subdomains". The Journal of Biological Chemistry. 280 (26): 24610–7. doi:10.1074/jbc.M413565200. PMID 15857831.
Further reading
- Kraft LJ, Kenworthy AK (January 2012). "Imaging protein complex formation in the autophagy pathway: analysis of the interaction of LC3 and Atg4B(C74A) in live cells using Förster resonance energy transfer and fluorescence recovery after photobleaching". Journal of Biomedical Optics. 17 (1): 011008. doi:10.1117/1.JBO.17.1.011008. PMC 3380812. PMID 22352642.
- Behrends C, Sowa ME, Gygi SP, Harper JW (July 2010). "Network organization of the human autophagy system". Nature. 466 (7302): 68–76. doi:10.1038/nature09204. PMC 2901998. PMID 20562859.
- Tanida I, Ueno T, Kominami E (December 2004). "LC3 conjugation system in mammalian autophagy". The International Journal of Biochemistry & Cell Biology. 36 (12): 2503–18. doi:10.1016/j.biocel.2004.05.009. PMID 15325588.
- Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T (November 2000). "LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing". The EMBO Journal. 19 (21): 5720–8. doi:10.1093/emboj/19.21.5720. PMC 305793. PMID 11060023.
- Tanida I, Tanida-Miyake E, Komatsu M, Ueno T, Kominami E (April 2002). "Human Apg3p/Aut1p homologue is an authentic E2 enzyme for multiple substrates, GATE-16, GABARAP, and MAP-LC3, and facilitates the conjugation of hApg12p to hApg5p". The Journal of Biological Chemistry. 277 (16): 13739–44. doi:10.1074/jbc.M200385200. PMID 11825910.
- Tanida I, Tanida-Miyake E, Nishitani T, Komatsu M, Yamazaki H, Ueno T, Kominami E (March 2002). "Murine Apg12p has a substrate preference for murine Apg7p over three Apg8p homologs". Biochemical and Biophysical Research Communications. 292 (1): 256–62. doi:10.1006/bbrc.2002.6645. PMID 11890701.
- He H, Dang Y, Dai F, Guo Z, Wu J, She X, Pei Y, Chen Y, Ling W, Wu C, Zhao S, Liu JO, Yu L (August 2003). "Post-translational modifications of three members of the human MAP1LC3 family and detection of a novel type of modification for MAP1LC3B". The Journal of Biological Chemistry. 278 (31): 29278–87. doi:10.1074/jbc.M303800200. PMID 12740394.
- Tanida I, Sou YS, Ezaki J, Minematsu-Ikeguchi N, Ueno T, Kominami E (August 2004). "HsAtg4B/HsApg4B/autophagin-1 cleaves the carboxyl termini of three human Atg8 homologues and delipidates microtubule-associated protein light chain 3- and GABAA receptor-associated protein-phospholipid conjugates". The Journal of Biological Chemistry. 279 (35): 36268–76. doi:10.1074/jbc.M401461200. PMID 15187094.
- Tanida I, Ueno T, Kominami E (November 2004). "Human light chain 3/MAP1LC3B is cleaved at its carboxyl-terminal Met121 to expose Gly120 for lipidation and targeting to autophagosomal membranes". The Journal of Biological Chemistry. 279 (46): 47704–10. doi:10.1074/jbc.M407016200. PMID 15355958.