Histone H2B type 2-E is a protein that in humans is encoded by the HIST2H2BE gene.[3][4][5]
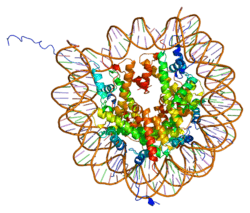
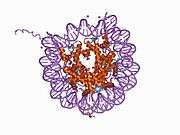
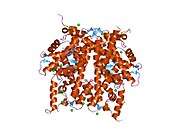
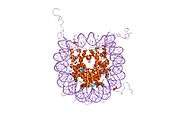
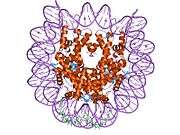
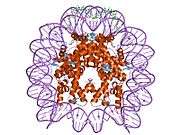
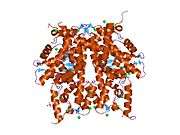
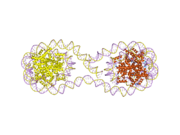
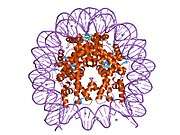
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. The linker histone, H1, interacts with linker DNA between nucleosomes and functions in the compaction of chromatin into higher order structures. This gene encodes a member of the histone H2B family, and generates two transcripts through the use of the conserved stem-loop termination motif, and the polyA addition motif.[5]
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000184678 - Ensembl, May 2017
- ↑ "Human PubMed Reference:".
- ↑ Collart D, Romain PL, Huebner K, Pockwinse S, Pilapil S, Cannizzaro LA, Lian JB, Croce CM, Stein JL, Stein GS (Jan 1993). "A human histone H2B.1 variant gene, located on chromosome 1, utilizes alternative 3' end processing". J Cell Biochem. 50 (4): 374–85. doi:10.1002/jcb.240500406. PMID 1469070.
- ↑ Marzluff WF, Gongidi P, Woods KR, Jin J, Maltais LJ (Oct 2002). "The human and mouse replication-dependent histone genes". Genomics. 80 (5): 487–98. doi:10.1016/S0888-7543(02)96850-3. PMID 12408966.
- 1 2 "Entrez Gene: HIST2H2BE histone cluster 2, H2be".
Further reading
- Collart D, Ramsey-Ewing A, Bortell R, et al. (1991). "Isolation and characterization of a cDNA from a human histone H2B gene which is reciprocally expressed in relation to replication-dependent H2B histone genes during HL60 cell differentiation". Biochemistry. 30 (6): 1610–7. doi:10.1021/bi00220a024. PMID 1993178.
- Jackson S, Brooks W, Jackson V (1994). "Dynamics of the interactions of histones H2A,H2B and H3,H4 with torsionally stressed DNA". Biochemistry. 33 (18): 5392–403. doi:10.1021/bi00184a006. PMID 8180162.
- Frohm M, Gunne H, Bergman AC, et al. (1996). "Biochemical and antibacterial analysis of human wound and blister fluid". Eur. J. Biochem. 237 (1): 86–92. doi:10.1111/j.1432-1033.1996.0086n.x. PMID 8620898.
- Rodriguez P, Munroe D, Prawitt D, et al. (1997). "Functional characterization of human nucleosome assembly protein-2 (NAP1L4) suggests a role as a histone chaperone". Genomics. 44 (3): 253–65. doi:10.1006/geno.1997.4868. PMID 9325046.
- El Kharroubi A, Piras G, Zensen R, Martin MA (1998). "Transcriptional Activation of the Integrated Chromatin-Associated Human Immunodeficiency Virus Type 1 Promoter". Mol. Cell. Biol. 18 (5): 2535–44. doi:10.1128/mcb.18.5.2535. PMC 110633. PMID 9566873.
- Zhang Y, Sun ZW, Iratni R, et al. (1998). "SAP30, a novel protein conserved between human and yeast, is a component of a histone deacetylase complex". Mol. Cell. 1 (7): 1021–31. doi:10.1016/S1097-2765(00)80102-1. PMID 9651585.
- Lorain S, Quivy JP, Monier-Gavelle F, et al. (1998). "Core Histones and HIRIP3, a Novel Histone-Binding Protein, Directly Interact with WD Repeat Protein HIRA". Mol. Cell. Biol. 18 (9): 5546–56. PMC 109139. PMID 9710638.
- Khan IU, Wallin R, Gupta RS, Kammer GM (1998). "Protein kinase A-catalyzed phosphorylation of heat shock protein 60 chaperone regulates its attachment to histone 2B in the T lymphocyte plasma membrane". Proc. Natl. Acad. Sci. U.S.A. 95 (18): 10425–30. doi:10.1073/pnas.95.18.10425. PMC 27910. PMID 9724719.
- Becker W, Weber Y, Wetzel K, et al. (1998). "Sequence characteristics, subcellular localization, and substrate specificity of DYRK-related kinases, a novel family of dual specificity protein kinases". J. Biol. Chem. 273 (40): 25893–902. doi:10.1074/jbc.273.40.25893. PMID 9748265.
- Allen MP, Zeng C, Schneider K, et al. (1999). "Growth arrest-specific gene 6 (Gas6)/adhesion related kinase (Ark) signaling promotes gonadotropin-releasing hormone neuronal survival via extracellular signal-regulated kinase (ERK) and Akt". Mol. Endocrinol. 13 (2): 191–201. doi:10.1210/me.13.2.191. PMID 9973250.
- Piredda L, Farrace MG, Lo Bello M, et al. (1999). "Identification of 'tissue' transglutaminase binding proteins in neural cells committed to apoptosis". FASEB J. 13 (2): 355–64. PMID 9973324.
- Carrier F, Georgel PT, Pourquier P, et al. (1999). "Gadd45, a p53-Responsive Stress Protein, Modifies DNA Accessibility on Damaged Chromatin". Mol. Cell. Biol. 19 (3): 1673–85. PMC 83961. PMID 10022855.
- Kawasaki H, Schiltz L, Chiu R, et al. (2000). "ATF-2 has intrinsic histone acetyltransferase activity which is modulated by phosphorylation". Nature. 405 (6783): 195–200. doi:10.1038/35012097. PMID 10821277.
- Deng L, de la Fuente C, Fu P, et al. (2001). "Acetylation of HIV-1 Tat by CBP/P300 increases transcription of integrated HIV-1 genome and enhances binding to core histones". Virology. 277 (2): 278–95. doi:10.1006/viro.2000.0593. PMID 11080476.
- Baake M, Doenecke D, Albig W (2001). "Characterisation of nuclear localisation signals of the four human core histones". J. Cell. Biochem. 81 (2): 333–46. doi:10.1002/1097-4644(20010501)81:2<333::AID-JCB1048>3.0.CO;2-D. PMID 11241673.
- Chadwick BP, Willard HF (2001). "A Novel Chromatin Protein, Distantly Related to Histone H2a, Is Largely Excluded from the Inactive X Chromosome". J. Cell Biol. 152 (2): 375–84. doi:10.1083/jcb.152.2.375. PMC 2199617. PMID 11266453.
- Freire J, Covelo G, Sarandeses C, et al. (2001). "Identification of nuclear-import and cell-cycle regulatory proteins that bind to prothymosin alpha". Biochem. Cell Biol. 79 (2): 123–31. doi:10.1139/bcb-79-2-123. PMID 11310559.
- Nemergut ME, Mizzen CA, Stukenberg T, et al. (2001). "Chromatin docking and exchange activity enhancement of RCC1 by histones H2A and H2B". Science. 292 (5521): 1540–3. doi:10.1126/science.292.5521.1540. PMID 11375490.
- Deng L, Wang D, de la Fuente C, et al. (2001). "Enhancement of the p300 HAT activity by HIV-1 Tat on chromatin DNA". Virology. 289 (2): 312–26. doi:10.1006/viro.2001.1129. PMID 11689053.
PDB gallery |
|---|
1aoi: COMPLEX BETWEEN NUCLEOSOME CORE PARTICLE (H3,H4,H2A,H2B) AND 146 BP LONG DNA FRAGMENT 1eqz: X-RAY STRUCTURE OF THE NUCLEOSOME CORE PARTICLE AT 2.5 A RESOLUTION 1f66: 2.6 A CRYSTAL STRUCTURE OF A NUCLEOSOME CORE PARTICLE CONTAINING THE VARIANT HISTONE H2A.Z 1hq3: CRYSTAL STRUCTURE OF THE HISTONE-CORE-OCTAMER IN KCL/PHOSPHATE 1kx3: X-Ray Structure of the Nucleosome Core Particle, NCP146, at 2.0 A Resolution 1kx4: X-Ray Structure of the Nucleosome Core Particle, NCP146b, at 2.6 A Resolution 1kx5: X-Ray Structure of the Nucleosome Core Particle, NCP147, at 1.9 A Resolution 1m18: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA 1m19: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA 1m1a: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA 1p34: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3a: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3b: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3f: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3g: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3i: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3k: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3l: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3m: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3o: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1p3p: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants 1s32: Molecular Recognition of the Nucleosomal 'Supergroove' 1tzy: Crystal Structure of the Core-Histone Octamer to 1.90 Angstrom Resolution 1u35: Crystal structure of the nucleosome core particle containing the histone domain of macroH2A 1zbb: Structure of the 4_601_167 Tetranucleosome 1zla: X-ray Structure of a Kaposi's sarcoma herpesvirus LANA peptide bound to the nucleosomal core 2aro: Crystal Structure Of The Native Histone Octamer To 2.1 Angstrom Resolution, Crystalised In The Presence Of S-Nitrosoglutathione 2cv5: Crystal structure of human nucleosome core particle 2f8n: 2.9 Angstrom X-ray structure of hybrid macroH2A nucleosomes 2fj7: Crystal structure of Nucleosome Core Particle Containing a Poly (dA.dT) Sequence Element 2hio: HISTONE OCTAMER (CHICKEN), CHROMOSOMAL PROTEIN 2nzd: Nucleosome core particle containing 145 bp of DNA |