ST turn
The ST turn is a structural feature in proteins and polypeptides.[1] Each consists of three amino acid residues (labeled i, i + 1 and i + 2) in which residue i is a serine (S) or threonine (T) that forms a hydrogen bond from its sidechain oxygen group to the mainchain NH group of residue i + 2.[2][3]
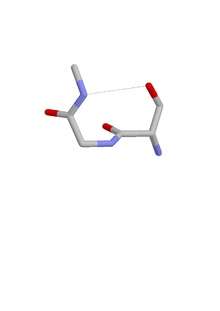
Similar motifs occur with aspartate or asparagine as residue i, called asx turn. Four types of asx turn and ST turn can be distinguished: types I, I’, II and II’. These categories correspond (via sidechain-mainchain mimicry of residue i) to those of the more abundant hydrogen-bonded beta turns, which have four residues and a hydrogen bond between the CO of residue i and the NH of residue i + 3. Regarding their occurrence in proteins, they differ in that type I is the commonest of the four beta turns while type II’ is the commonest of the ST and asx turns.
Asx and ST turns both occur frequently at the N-termini of α-helices,[4][5][6][7] as part of asx motifs or ST motifs, with the asx, serine or threonine as the N cap residue. They are thus often regarded as helix capping features.
Evidence for a functionally relevant ST turn is provided in the CDR3 region of the T-cell receptor (B chain, V domain) [8]
A proportion of ST turns are accompanied by a mainchain-mainchain hydrogen bond that qualifies them as ST motifs.
References
- Duddy, WJ; Nissink WMJ; Allen, Frank H.; Milner-White, E. James (2004). "Mimicry by asx- and ST-turns of the four main types of β-turn in proteins". Protein Science. 13 (11): 3051–3055. doi:10.1110/ps.04920904. PMC 2286581. PMID 15459339.
- Leader, DP; Milner-White EJ (2009). "Motivated Proteins: A web application for studying small three-dimensional protein motifs". BMC Bioinformatics. 10: 60. doi:10.1186/1471-2105-10-60. PMC 2651126. PMID 19210785.
- Golovin, A; Henrick K (2008). "MSDmotif: exploring protein sites and motifs". BMC Bioinformatics. 9: 312. doi:10.1186/1471-2105-9-312. PMC 2491636. PMID 18637174.
- Doig, AJ; Macarthur MW; MacArthur, Malcolm W.; Thornton, Janet M. (2008). "Structures of N-termini of helices in proteins". Protein Science. 6 (1): 147–155. doi:10.1002/pro.5560060117. PMC 2143508. PMID 9007987.
- Presta, LG; Rose GD (1988). "Helix Caps". Science. 240 (4859): 1632–1641. Bibcode:1988Sci...240.1632P. doi:10.1126/science.2837824. PMID 2837824.
- Aurora, R; Rose GD (1998). "Helix Capping". Protein Science. 7 (1): 21–38. doi:10.1002/pro.5560070103. PMC 2143812. PMID 9514257.
- Gunasekaran, K; Nagarajam HA; Ramakrishnan, C; Balaram, P (1998). "Sterochemical punctuation marks in protein structure". Journal of Molecular Biology. 275 (5): 917–932. doi:10.1006/jmbi.1997.1505. PMID 9480777.
- Yassai, MB; Demus W; Gorski J (2017). "Structural and Mechanistic Implications of Rearrangement Frequencies within Human TCRBV Genes". J Immunol. 199 (3): 1142–1152. doi:10.4049/jimmunol.1601450. PMC 5659713. PMID 28659354.